|
(1993) Optimal alignment between groups of sequences and its application to multiple sequence alignment. (1991) A novel randomized iterative strategy for aligning multiple protein sequences. 
confidence levels from tertiary structure comparisons. (1987) A strategy for the rapid multiple alignment of protein sequences. 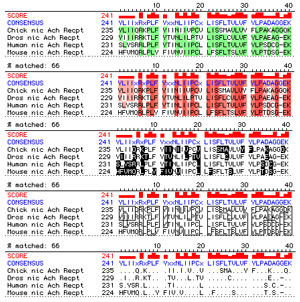
(2007) Parttree: an algorithm to build an approximate tree from a large number of unaligned sequences. (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. (1987) Progressive sequence alignment as a prerequisite to correct phylogenetic trees. (1982) An improved algorithm for matching biological sequences. (1981) Identification of common molecular subsequences. (1970) A general method applicable to the search for similarities in the amino acid sequence of two proteins. (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. (2004) Aligning multiple genomic sequences with the threaded blockset aligner. M., Baertsch, R., Rosenbloom, K., Clawson, H., Green, E. (2007) DNA reference alignment benchmarks based on tertiary structure of encoded proteins. Algorithms Mol Biol 1, 19.Ĭarroll, H., Beckstead, W., O’connor, T., Ebbert, M., Clement, M., Snell, Q., and McClellan, D. (2006) An enhanced RNA alignment benchmark for sequence alignment programs. (2005) MAFFT version 5: improvement in accuracy of multiple sequence alignment. Katoh, K., Kuma, K., Toh, H., and Miyata, T. (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Katoh, K., Misawa, K., Kuma, K., and Miyata, T. (2003) Leveraging the mouse genome for gene prediction in human: from whole-genome shotgun reads to a global synteny map. Proc Natl Acad Sci USA 74, 5088–90.įlicek, P., Keibler, E., Hu, P., Korf, I., and Brent, M. (1977) Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Read our Privacy Notice if you are concerned with your privacy and how we handle personal information.Woese, C. If you plan to use these services during a course please contact us. 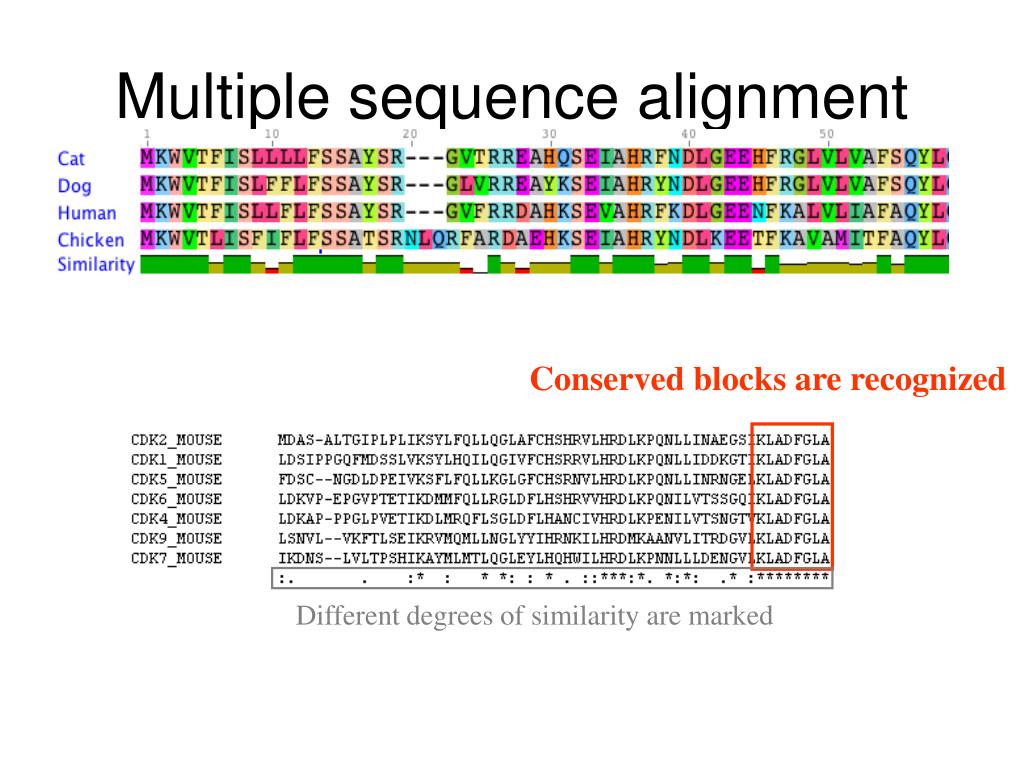
If you have any feedback or encountered any issues please let us know via EMBL-EBI Support. Please read the provided Help & Documentation and FAQs before seeking help from our support staff. 
The tools described on this page are provided using Search and sequence analysis tools services from EMBL-EBI in 2022 The EBI has a new phylogeny-aware multiple sequence alignment program which makes use of evolutionary information to help place insertions and deletions. Transform a Sequence Similarity Search result into a Multiple Sequence Alignment or reformat a Multiple Sequence Alignment using the MView program.Ĭonsistency-based MSA tool that attempts to mitigate the pitfalls of progressive alignment methods. Suitable for medium-large alignments.Īccurate MSA tool, especially good with proteins. MSA tool that uses Fast Fourier Transforms. Very fast MSA tool that concentrates on local regions. Suitable for medium-large alignments.ĮMBOSS Cons creates a consensus sequence from a protein or nucleotide multiple alignment. New MSA tool that uses seeded guide trees and HMM profile-profile techniques to generate alignments. From the output, homology can be inferred and the evolutionary relationships between the sequences studied.īy contrast, Pairwise Sequence Alignment tools are used to identify regions of similarity that may indicate functional, structural and/or evolutionary relationships between two biological sequences. Multiple Sequence Alignment (MSA) is generally the alignment of three or more biological sequences (protein or nucleic acid) of similar length.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
